Home > Press > A new single-molecule tool to observe enzymes at work
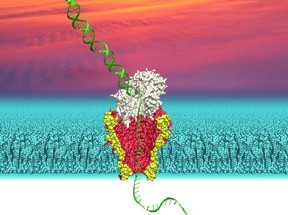 |
| This is an illustration of a nanopore derived from a genetically modified bacterial membrane channel with DNA passing through it. CREDIT: Ian Derrington, University of Washington |
Abstract:
A team of scientists at the University of Washington and the biotechnology company Illumina have created an innovative tool to directly detect the delicate, single-molecule interactions between DNA and enzymatic proteins. Their approach provides a new platform to view and record these nanoscale interactions in real time. As they report Sept. 28 in Nature Biotechnology, this tool should provide fast and reliable characterization of the different mechanisms cellular proteins use to bind to DNA strands -- information that could shed new light on the atomic-scale interactions within our cells and help design new drug therapies against pathogens by targeting enzymes that interact with DNA.
A new single-molecule tool to observe enzymes at work
Seattle, WA | Posted on September 28th, 2015"There are other single-molecule tools around, but our new tool is far more sensitive," said senior author and UW physics professor Jens Gundlach. "We can really pick up atomic-scale movements that a protein imparts onto DNA."
As can happen in the scientific process, they developed this tool -- the single-molecule picometer-resolution nanopore tweezers, or SPRNT -- while working on a related project.
The UW team has been exploring nanopore technology to read DNA sequences quickly. Our genes are long stretches of DNA molecules, which are made up of combinations of four chemical DNA "letters." In their approach, Gundlach and his team measure an electrical current through a biological pore called MspA, which is embedded within a modified cell membrane. As DNA passes through a tiny opening in the pore -- an opening that is just 0.00000012 centimeters wide, or 1/10,000th the width of a human hair -- the current shifts based on the sequence of DNA letters. They use these changes in current to infer DNA sequences.
Gundlach and his team, in the process of investigating nanopore sequencing, tried out a variety of molecular motors to move DNA through the pore. They discovered that their experimental setup was sensitive enough to observe motions much smaller than the distance between adjacent letters on the DNA. As they report in their paper, SPRNT is more than seven times more sensitive than existing techniques to measure interactions between DNA and proteins.
"Generally, most existing techniques to look at single-molecule movements -- such as optical tweezers -- have a resolution, at best, of about 300 picometers," said Gundlach. "With SPRNT, we can have 40 picometer resolution."
For reference, 40 picometers are 0.000000004 centimeters, or about 0.0000000016 inches.
"We realized we can detect minute differences in the position of the DNA in the pore," said UW physics postdoctoral researcher Andrew Laszlo, a co-author on the paper. "We could pick up differences in how the proteins were binding to DNA and moving it through the pore."
These differences account for the unique role each cellular protein plays as it interacts with DNA. Cells have proteins to copy DNA, "read" DNA to express genes and repair DNA when it is damaged. There are cellular proteins that unwind DNA, while others bunch DNA tightly together.
Biologists have long recognized that proteins have different structures to perform these roles, but the physical motion of proteins as they work on DNA has been difficult to detect directly.
"When you have the kind of resolution that SPRNT offers, you can start to pick apart the minute steps these proteins take," said Laszlo.
Gundlach and his team show that SPRNT is sensitive enough to differentiate between the mechanisms that two cellular proteins use to pass DNA through the nanopore opening. One protein, which normally copies DNA, moves along the DNA one letter at a time as it guides DNA through the pore. The second protein, which normally unwinds DNA, instead takes two steps along each DNA letter, which they could pick up by tracking minute changes in the current, according to co-author and UW physics doctoral student Jonathan Craig. They even discovered that these two steps involve sequential chemical processes that the protein uses to walk along DNA.
"You can really see the underlying mechanisms, and that has a ton of implications -- from understanding how life works to drug design," said Laszlo.
Gundlach believes this tool may open a new window for understanding how cellular proteins process DNA, which could help genetically engineer proteins to perform novel jobs. These fine details may also help scientists understand how mutations in proteins can lead to disease or find protein properties that would be ideal targets for drug therapies.
"For example, viral genes code for their own proteins that process their DNA," said Gundlach. "If we can use SPRNT to screen for drugs that specifically disrupt the functioning of these proteins, it may be possible to interfere with viruses."
###
Other UW authors on the paper include lead author Ian Derrington, a postdoctoral researcher in physics, and Brian Ross, Henry Brinkerhoff, Ian Nova, Kenji Doering and Benjamin Tickman. Co-authors at Illumina are Kevin Gunderson, Eric Stava, Mostafa Ronaghi and Jeffrey Mandell. Gundlach's laboratory received funding for this project from the National Institutes of Health's $1,000 Genome Project, grant number R01HG005115.
####
For more information, please click here
Contacts:
James Urton
206-543-2580
Jens Gundlach
206-543-8774
Copyright © University of Washington
If you have a comment, please Contact us.Issuers of news releases, not 7th Wave, Inc. or Nanotechnology Now, are solely responsible for the accuracy of the content.
| Related News Press |
News and information
![]() Simulating magnetization in a Heisenberg quantum spin chain April 5th, 2024
Simulating magnetization in a Heisenberg quantum spin chain April 5th, 2024
![]() NRL charters Navy�s quantum inertial navigation path to reduce drift April 5th, 2024
NRL charters Navy�s quantum inertial navigation path to reduce drift April 5th, 2024
![]() Discovery points path to flash-like memory for storing qubits: Rice find could hasten development of nonvolatile quantum memory April 5th, 2024
Discovery points path to flash-like memory for storing qubits: Rice find could hasten development of nonvolatile quantum memory April 5th, 2024
Govt.-Legislation/Regulation/Funding/Policy
![]() NRL charters Navy�s quantum inertial navigation path to reduce drift April 5th, 2024
NRL charters Navy�s quantum inertial navigation path to reduce drift April 5th, 2024
![]() Discovery points path to flash-like memory for storing qubits: Rice find could hasten development of nonvolatile quantum memory April 5th, 2024
Discovery points path to flash-like memory for storing qubits: Rice find could hasten development of nonvolatile quantum memory April 5th, 2024
![]() Chemical reactions can scramble quantum information as well as black holes April 5th, 2024
Chemical reactions can scramble quantum information as well as black holes April 5th, 2024
Nanomedicine
![]() New micromaterial releases nanoparticles that selectively destroy cancer cells April 5th, 2024
New micromaterial releases nanoparticles that selectively destroy cancer cells April 5th, 2024
![]() Good as gold - improving infectious disease testing with gold nanoparticles April 5th, 2024
Good as gold - improving infectious disease testing with gold nanoparticles April 5th, 2024
![]() Researchers develop artificial building blocks of life March 8th, 2024
Researchers develop artificial building blocks of life March 8th, 2024
Discoveries
![]() Chemical reactions can scramble quantum information as well as black holes April 5th, 2024
Chemical reactions can scramble quantum information as well as black holes April 5th, 2024
![]() New micromaterial releases nanoparticles that selectively destroy cancer cells April 5th, 2024
New micromaterial releases nanoparticles that selectively destroy cancer cells April 5th, 2024
![]() Utilizing palladium for addressing contact issues of buried oxide thin film transistors April 5th, 2024
Utilizing palladium for addressing contact issues of buried oxide thin film transistors April 5th, 2024
Announcements
![]() NRL charters Navy�s quantum inertial navigation path to reduce drift April 5th, 2024
NRL charters Navy�s quantum inertial navigation path to reduce drift April 5th, 2024
![]() Discovery points path to flash-like memory for storing qubits: Rice find could hasten development of nonvolatile quantum memory April 5th, 2024
Discovery points path to flash-like memory for storing qubits: Rice find could hasten development of nonvolatile quantum memory April 5th, 2024
Interviews/Book Reviews/Essays/Reports/Podcasts/Journals/White papers/Posters
![]() Simulating magnetization in a Heisenberg quantum spin chain April 5th, 2024
Simulating magnetization in a Heisenberg quantum spin chain April 5th, 2024
![]() Discovery points path to flash-like memory for storing qubits: Rice find could hasten development of nonvolatile quantum memory April 5th, 2024
Discovery points path to flash-like memory for storing qubits: Rice find could hasten development of nonvolatile quantum memory April 5th, 2024
Tools
![]() Ferroelectrically modulate the Fermi level of graphene oxide to enhance SERS response November 3rd, 2023
Ferroelectrically modulate the Fermi level of graphene oxide to enhance SERS response November 3rd, 2023
![]() The USTC realizes In situ electron paramagnetic resonance spectroscopy using single nanodiamond sensors November 3rd, 2023
The USTC realizes In situ electron paramagnetic resonance spectroscopy using single nanodiamond sensors November 3rd, 2023
Nanobiotechnology
![]() New micromaterial releases nanoparticles that selectively destroy cancer cells April 5th, 2024
New micromaterial releases nanoparticles that selectively destroy cancer cells April 5th, 2024
![]() Good as gold - improving infectious disease testing with gold nanoparticles April 5th, 2024
Good as gold - improving infectious disease testing with gold nanoparticles April 5th, 2024
![]() Researchers develop artificial building blocks of life March 8th, 2024
Researchers develop artificial building blocks of life March 8th, 2024
|
|
||
|
|
||
| The latest news from around the world, FREE | ||
|
|
||
|
|
||
| Premium Products | ||
|
|
||
|
Only the news you want to read!
Learn More |
||
|
|
||
|
Full-service, expert consulting
Learn More |
||
|
|
||








